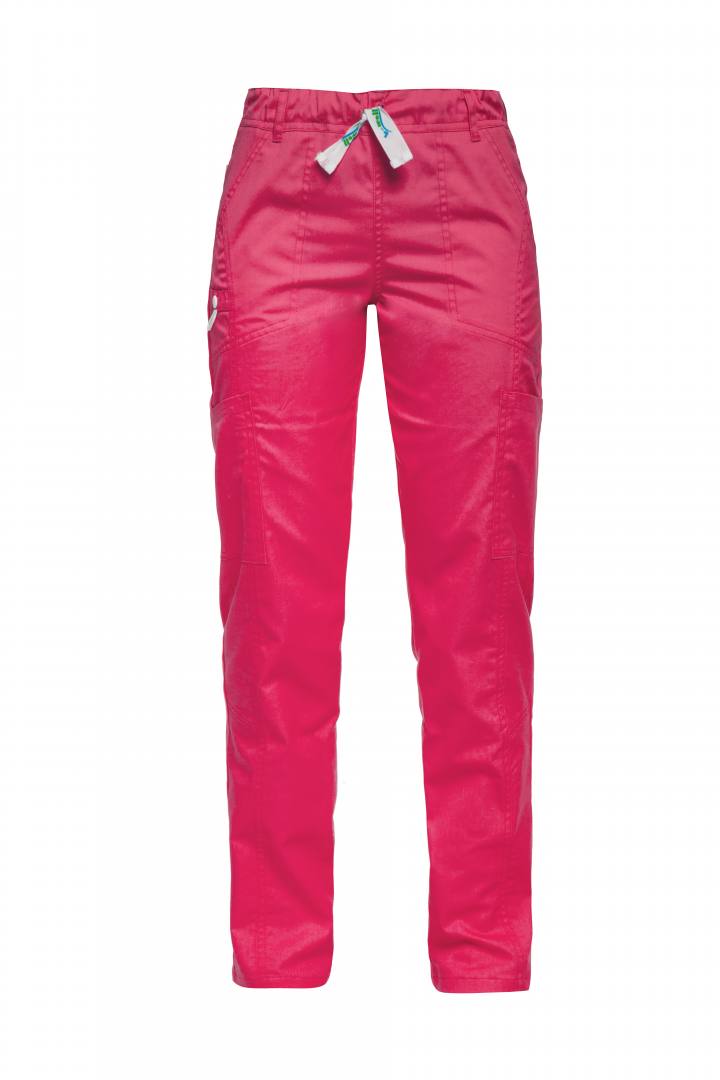
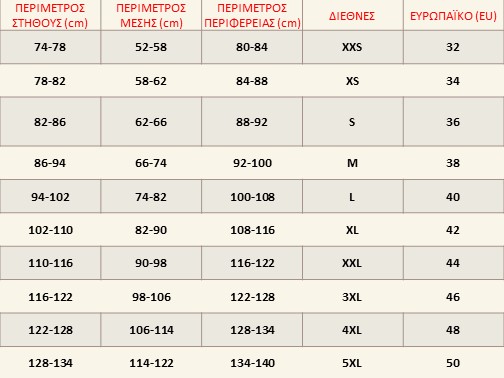
Παντελόνι unisex medical ΦΟΥΞΙΑ DANTE 180 gr/m² B-WELL εργασιας - WorkProtection Ένδυση Εργασίας Ρούχα εργασίας
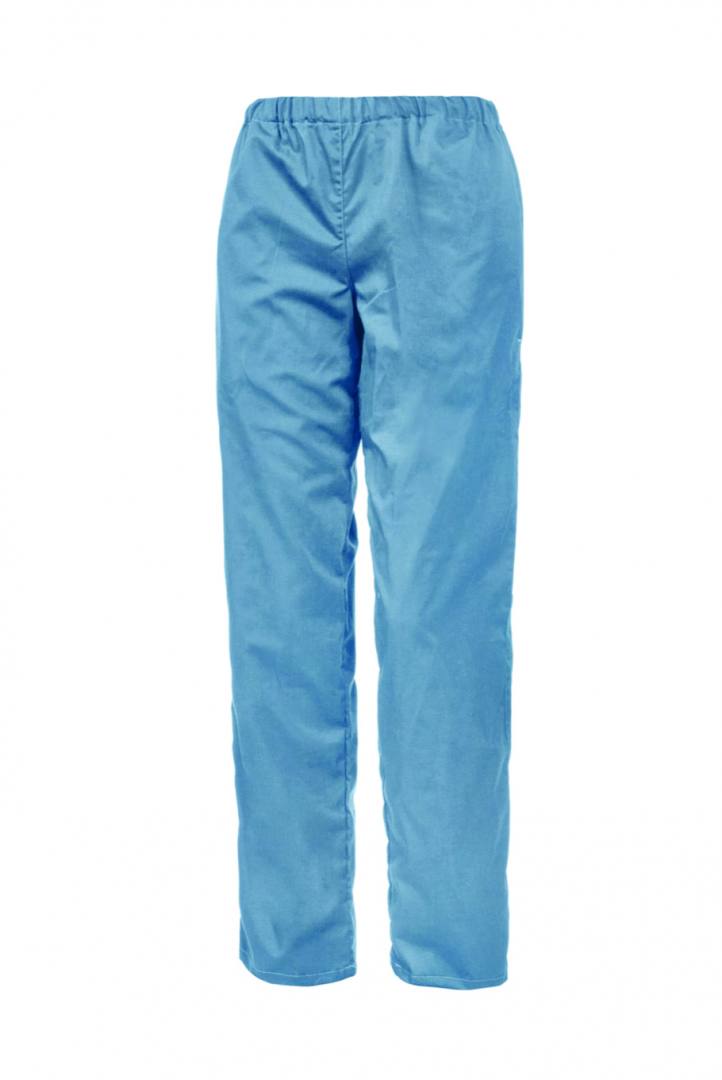

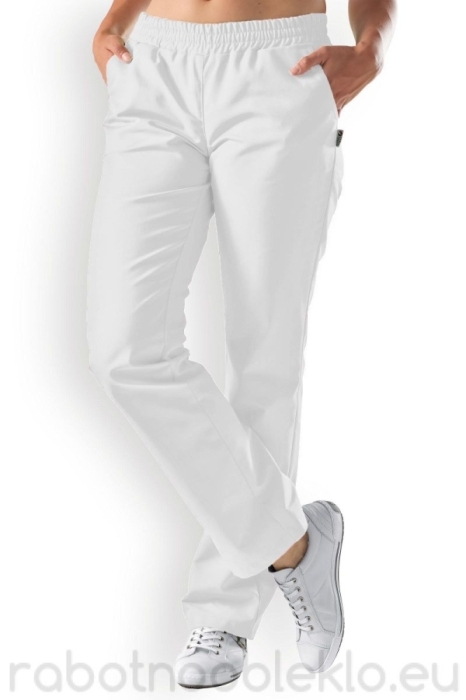
Παντελόνια unisex. BATISTA medical γαλάζιο 160 gr/m² B-WELL εργασιας - WorkProtection Ένδυση Εργασίας Ρούχα εργασίας
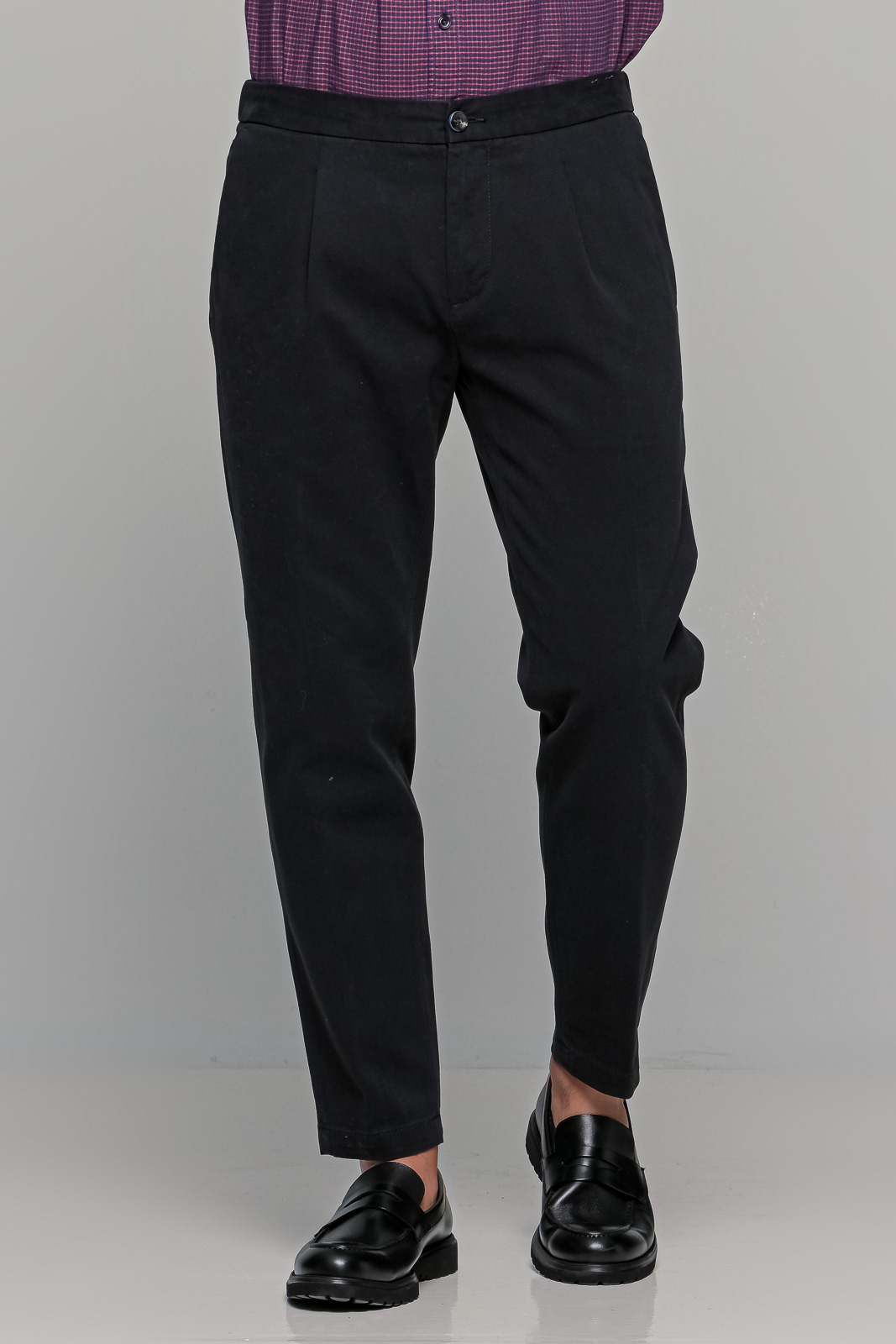

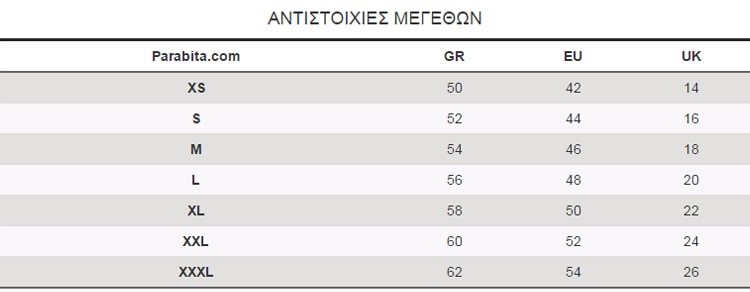
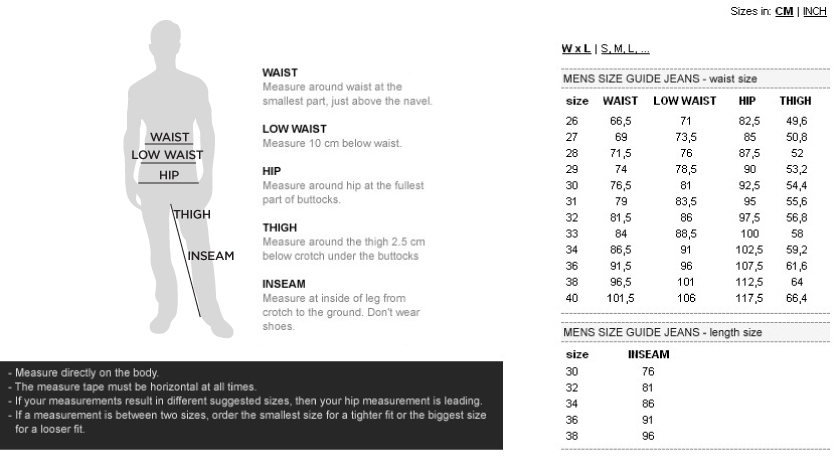
AT.P.CO. βαμβακερό ανδρικό παντελόνι Jogger - A251BERLINO563999 Μαύρο με €90,30 | Ανδρικά Ρούχα | TsitsoStore

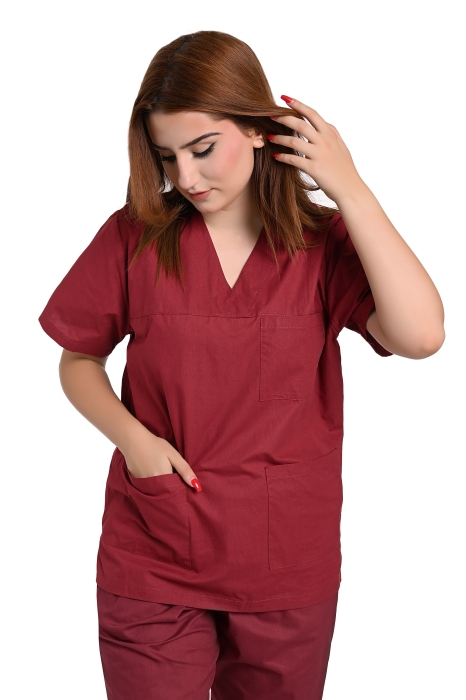
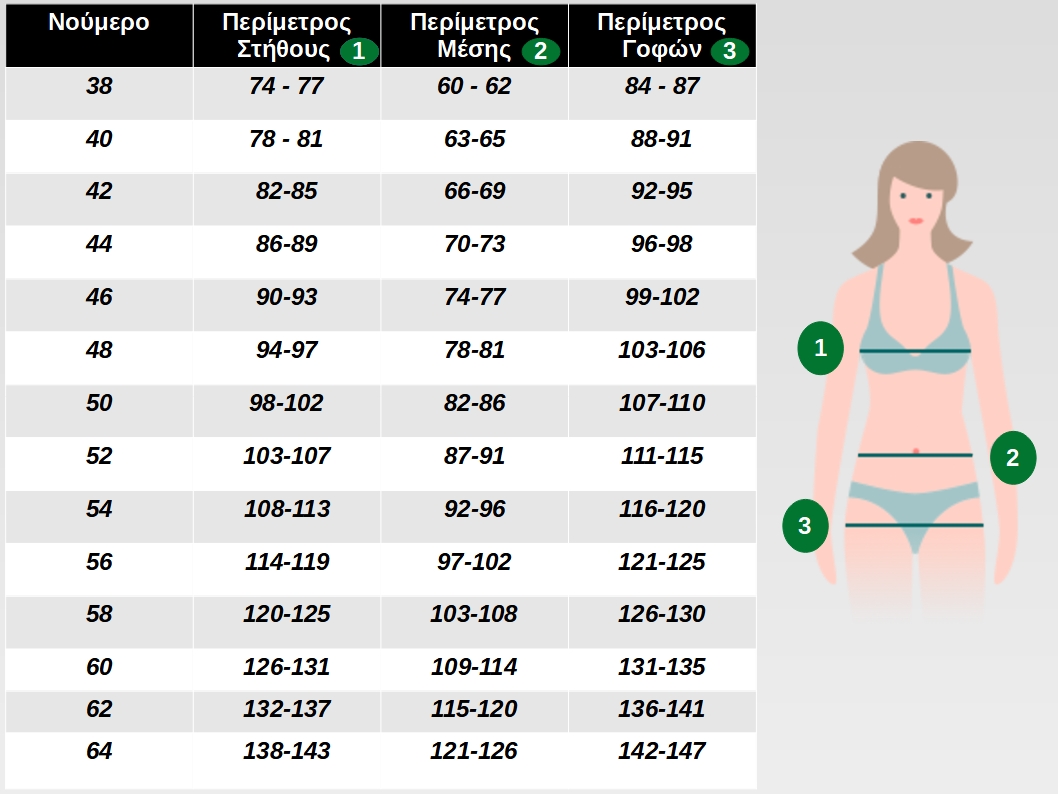
ιατρική νοσηλευτική μπλούζα γυναικεία medical BAMBINA Frogs B-WELL εργασιας - WorkProtection Ένδυση Εργασίας Ρούχα εργασίας

Παντελόνια unisex. BATISTA medical σκούρο μπλε 160 gr/m² B-WELL εργασιας - WorkProtection Ένδυση Εργασίας Ρούχα εργασίας