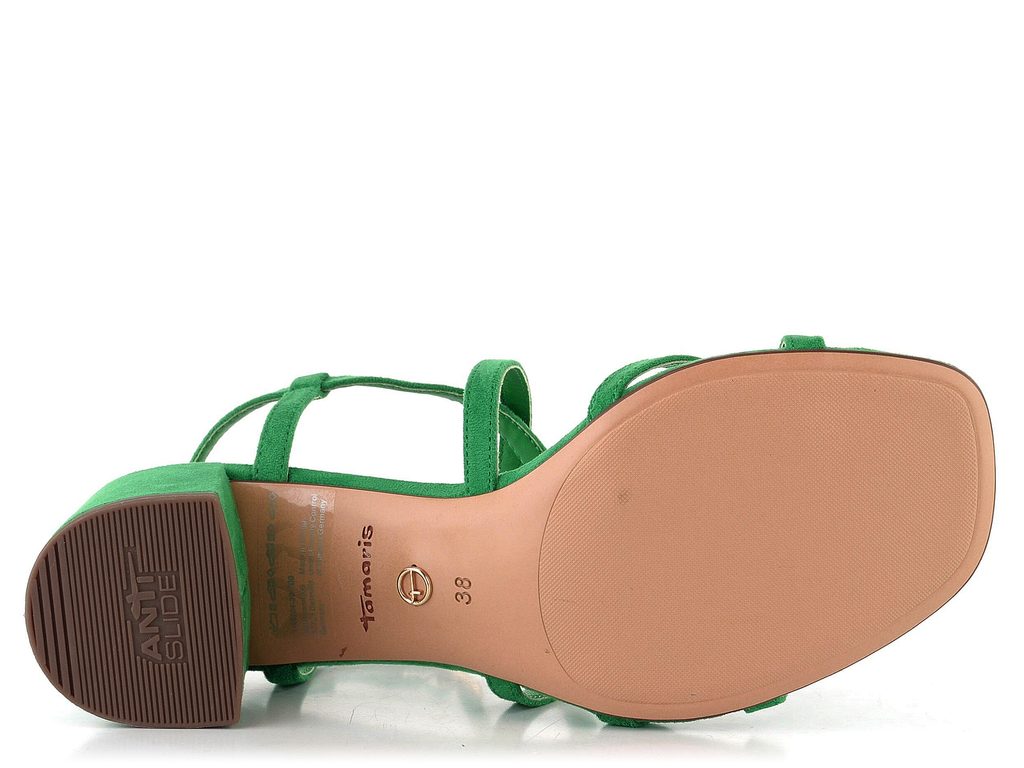

Tamaris zelené páskové sandálky na podpatku Green 1-28204-20 - Tamaris - Sandály - JADI.cz - ...více než boty
caliparks.org/app/public/assets/data/parks.json at parksforward-redesign · GreenInfo-Network/caliparks.org · GitHub
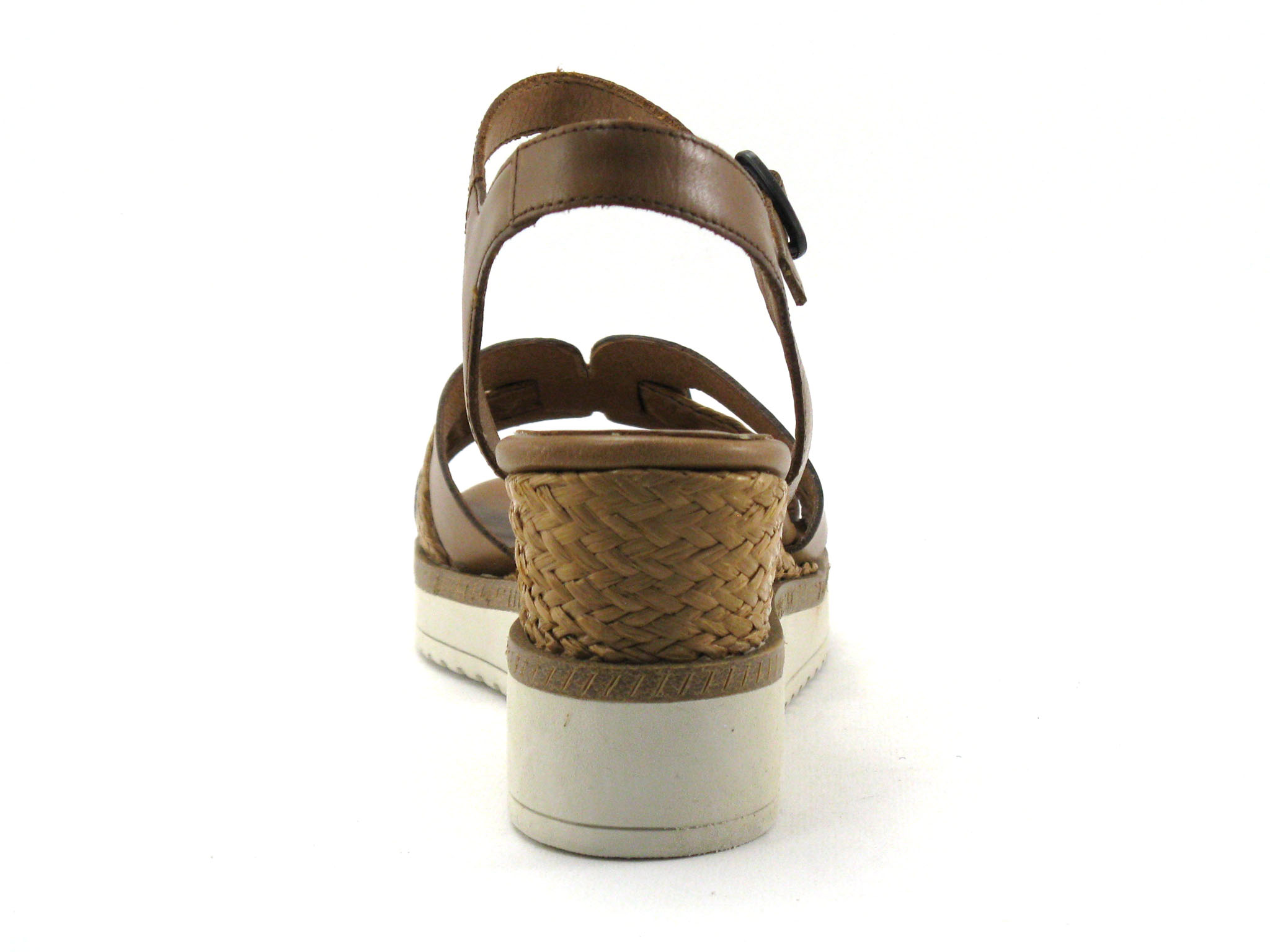

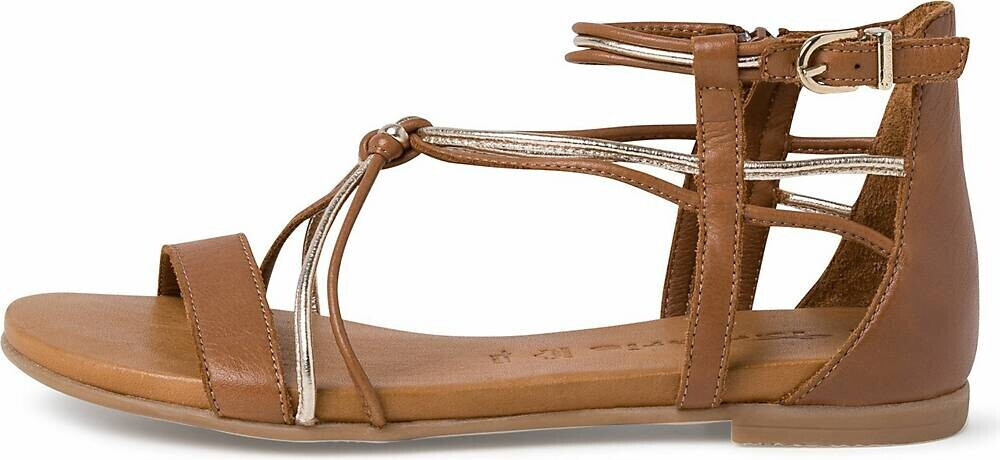
Achat chaussures Tamaris Femme Sandale et Nu-pieds, vente Tamaris 1-28243-28 Cognac - Nu-Pieds marron Femme - Talon compense
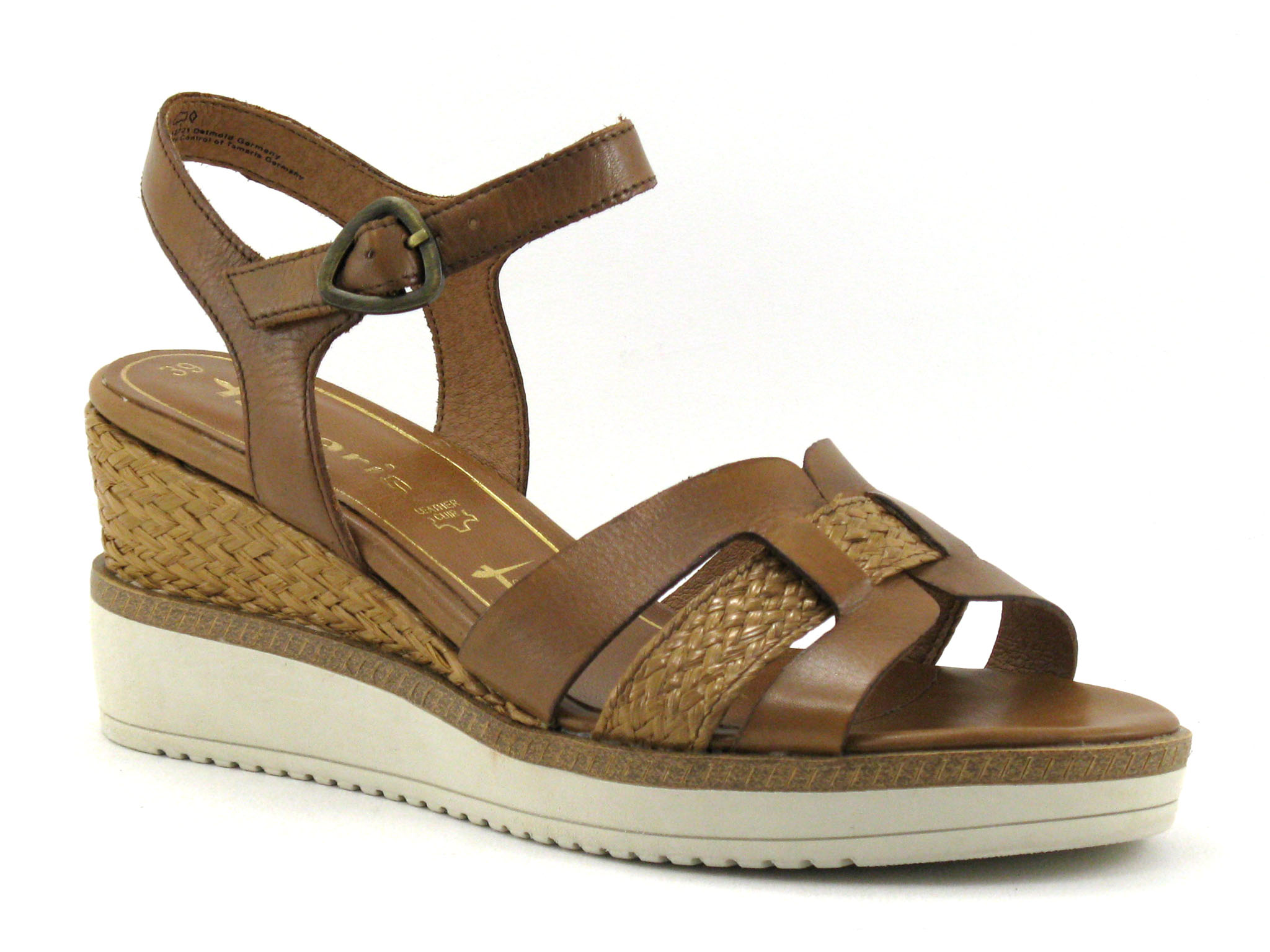

Achat chaussures Tamaris Femme Sandale et Nu-pieds, vente Tamaris 1-28243-28 Cognac - Nu-Pieds marron Femme - Talon compense
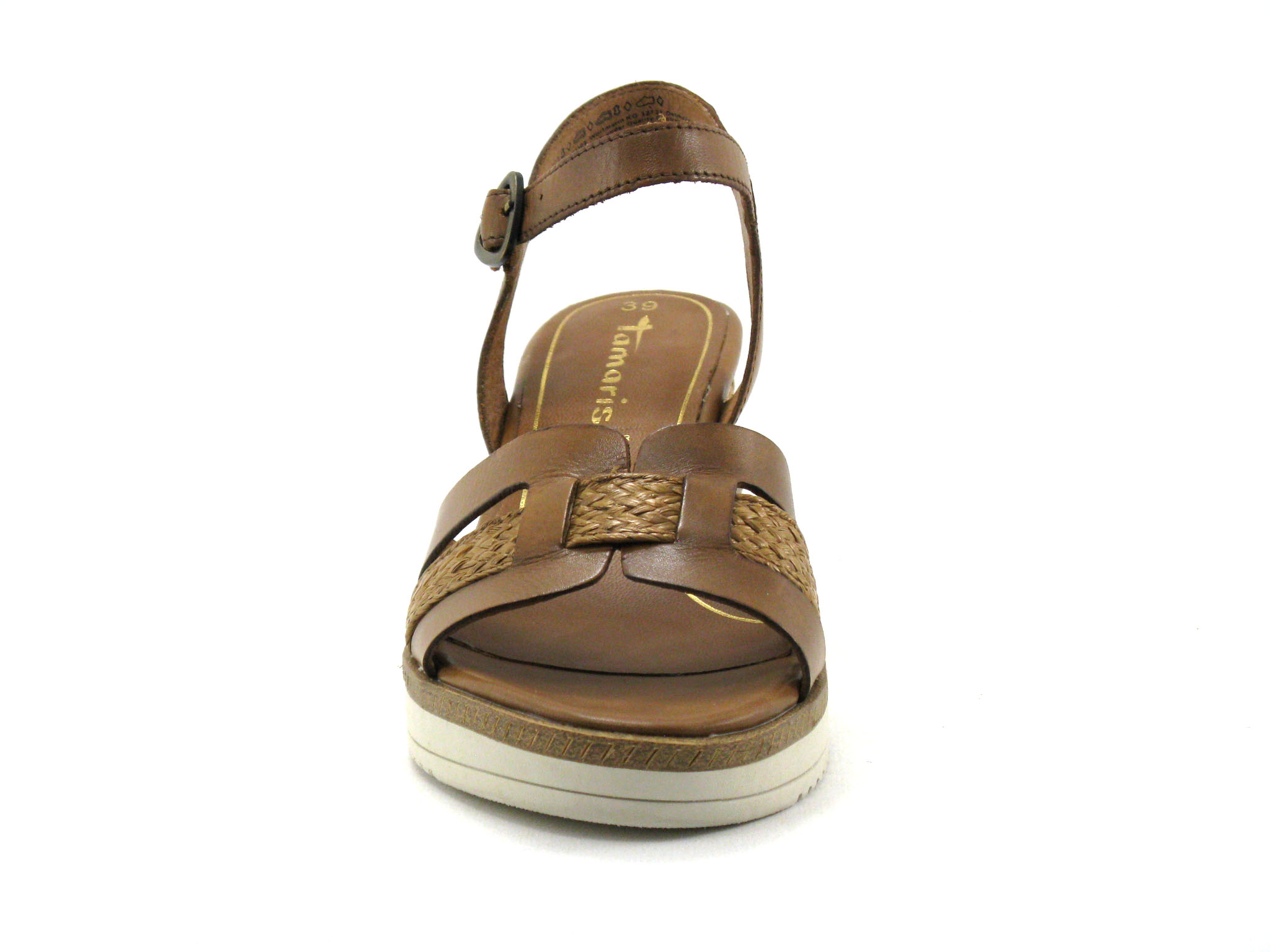
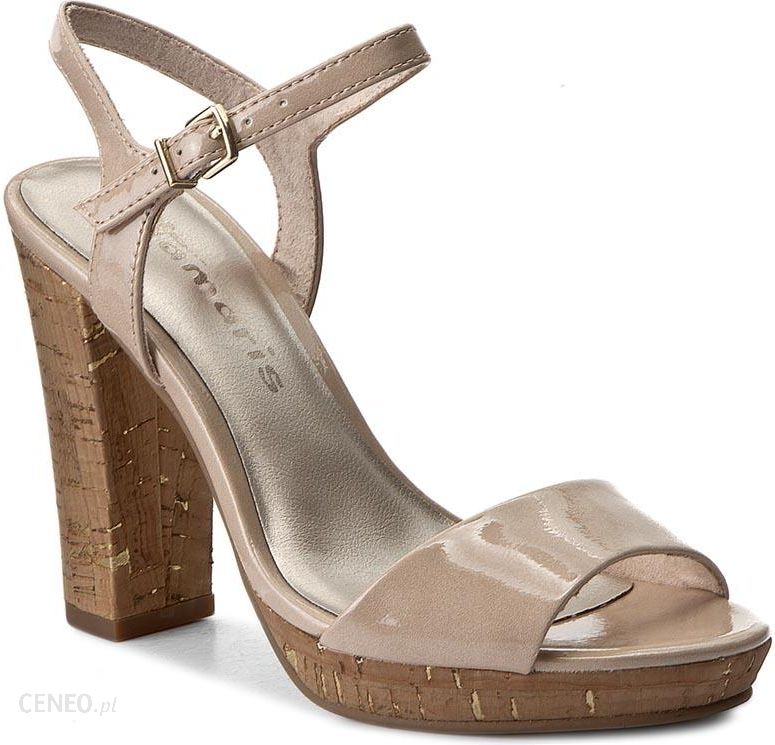
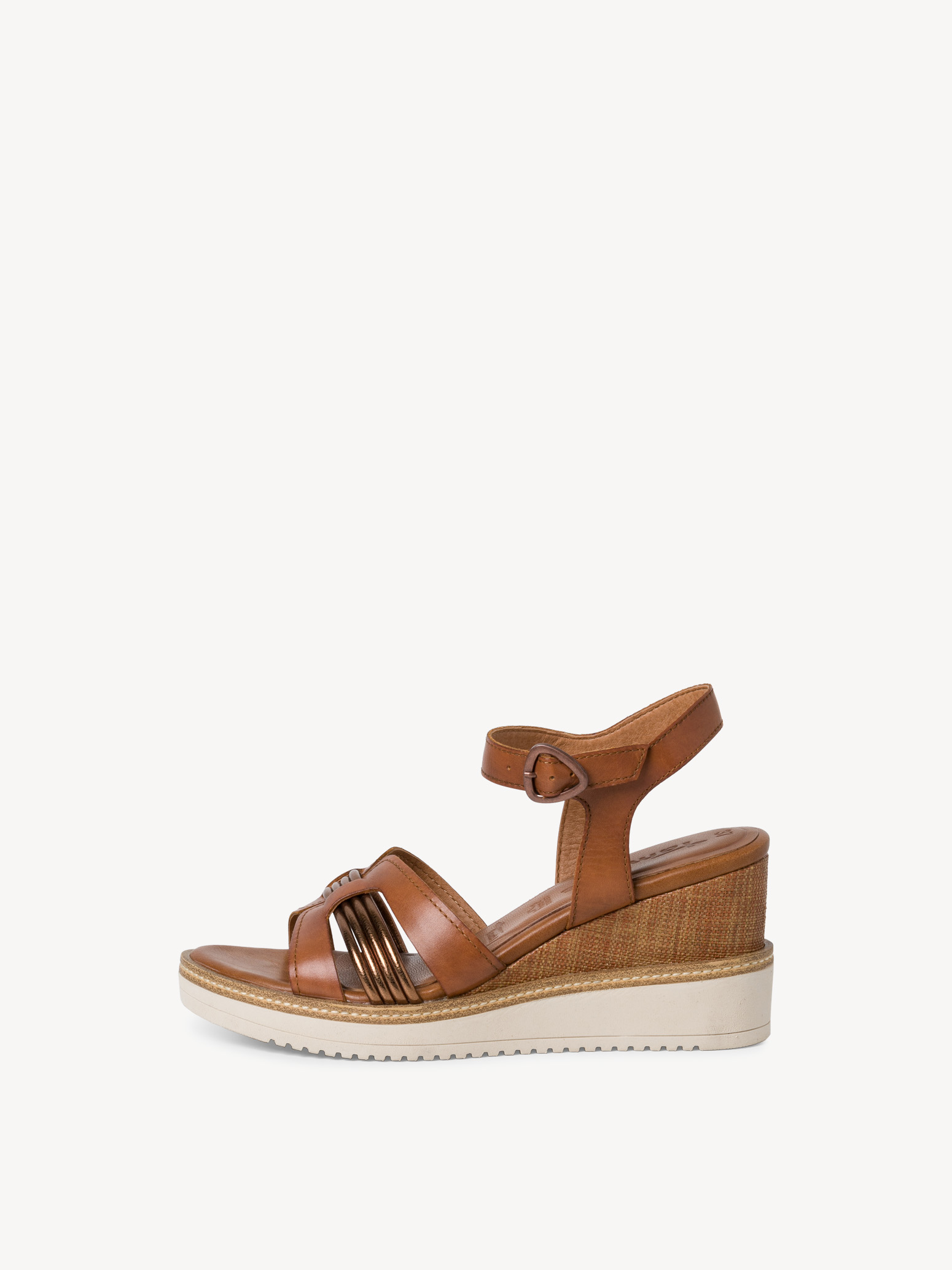
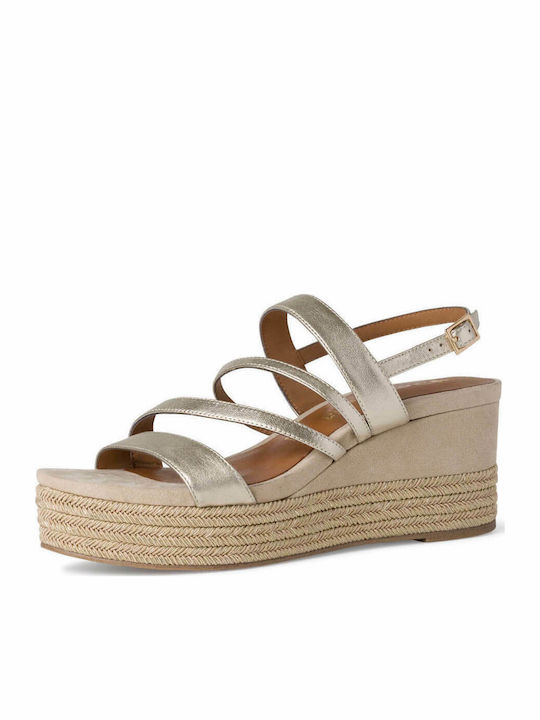
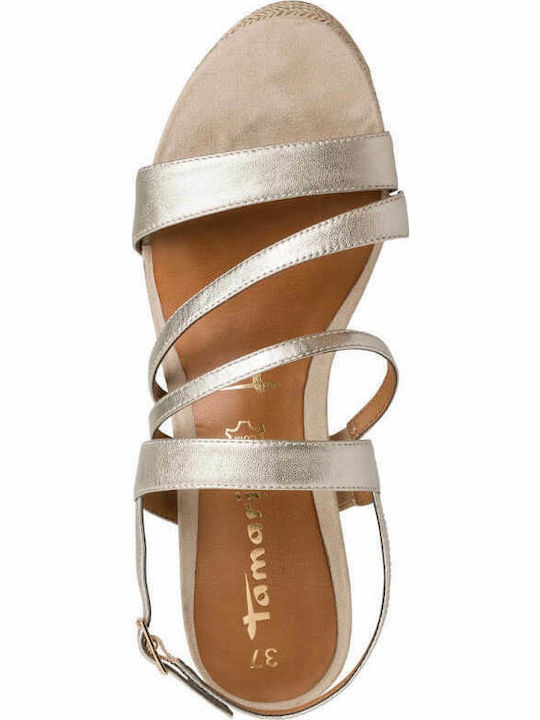
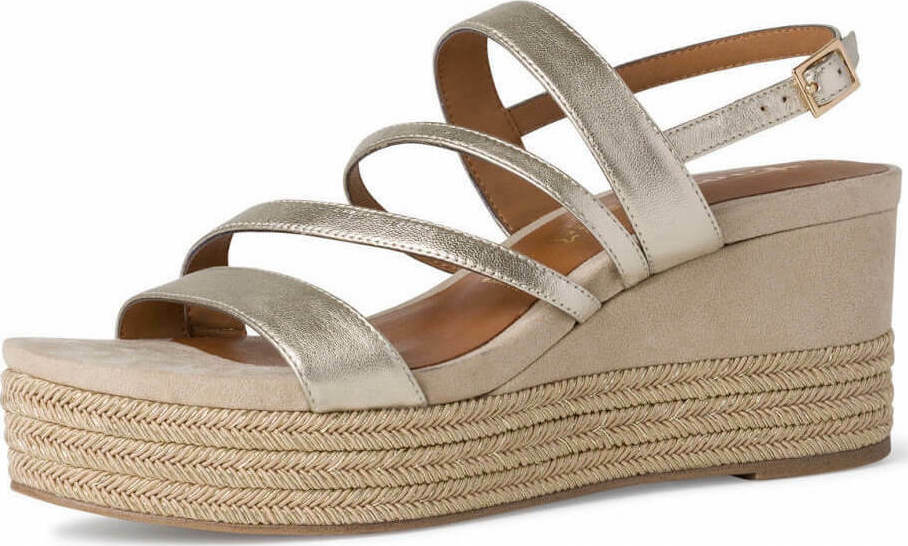
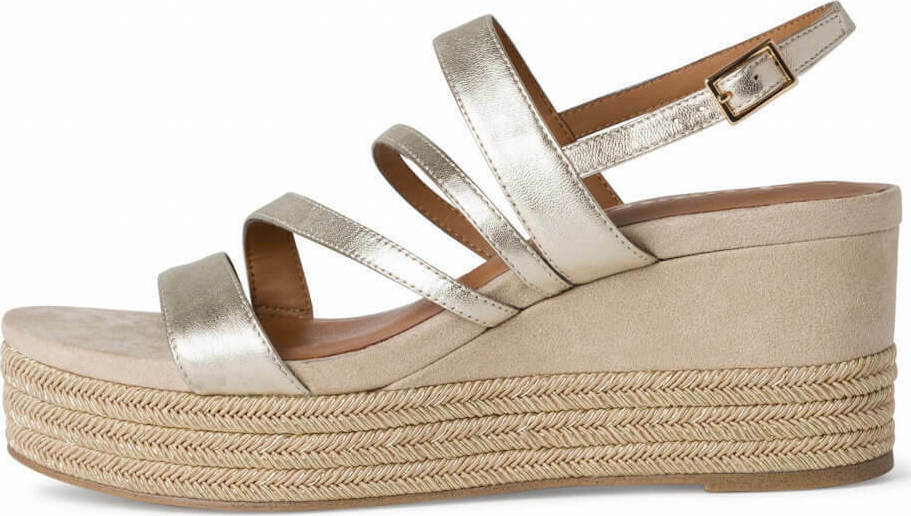
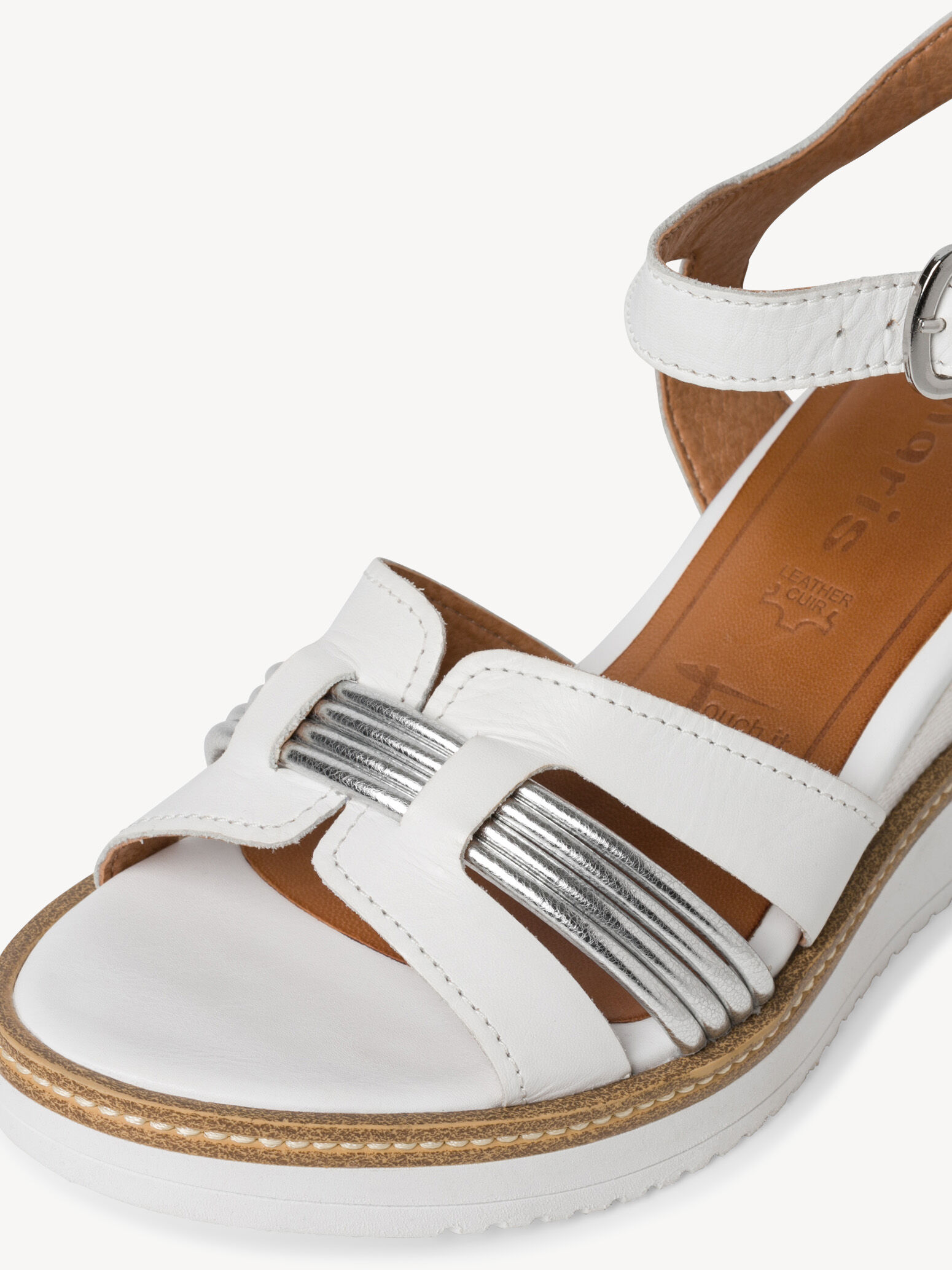
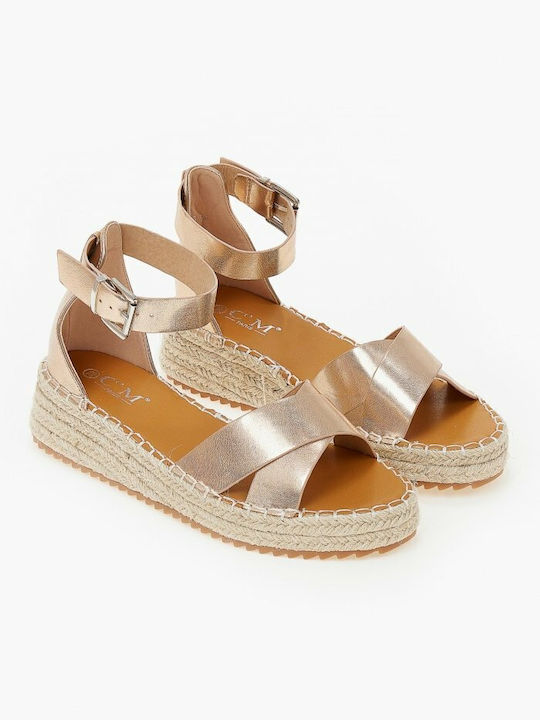














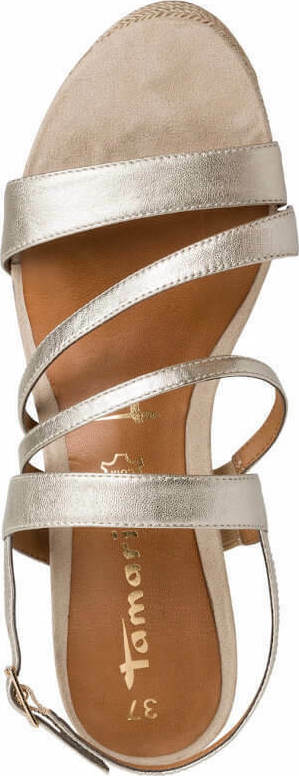









/szandal-tamaris-1-28281-38-beige-nu-gold-492.jpg)
