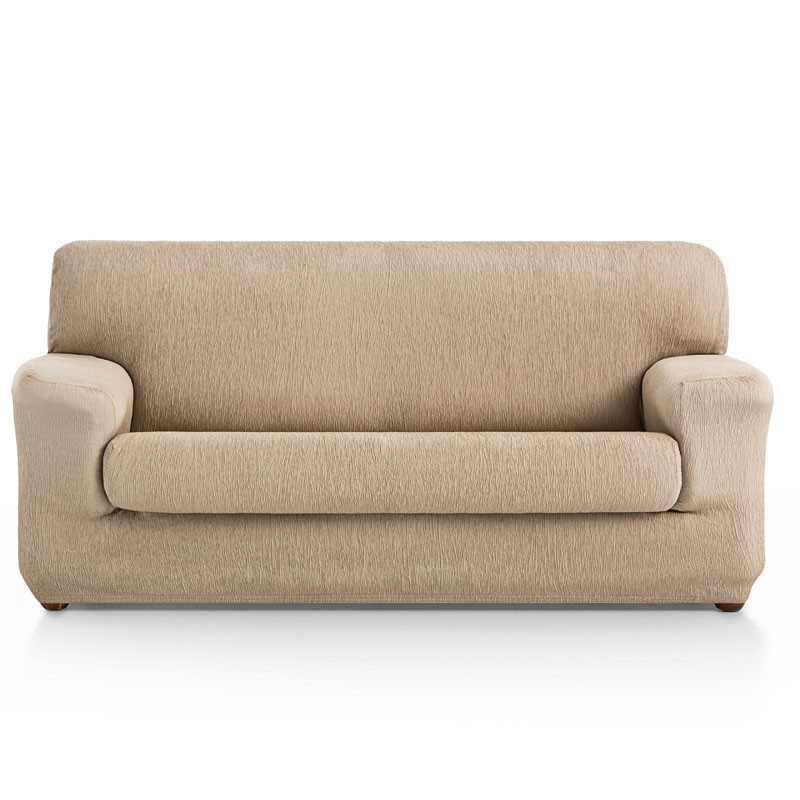
Ανταλλακτικό κάλυμμα μαξιλαριού καναπέ Ηρα - Εισαγωγή & εμπόριο προϊόντων εμποτισμένης ξυλείας για εξωτερική χρήση.

Ανταλλακτικό κάλυμμα μαξιλαριού καναπέ Ηρα - Εισαγωγή & εμπόριο προϊόντων εμποτισμένης ξυλείας για εξωτερική χρήση.

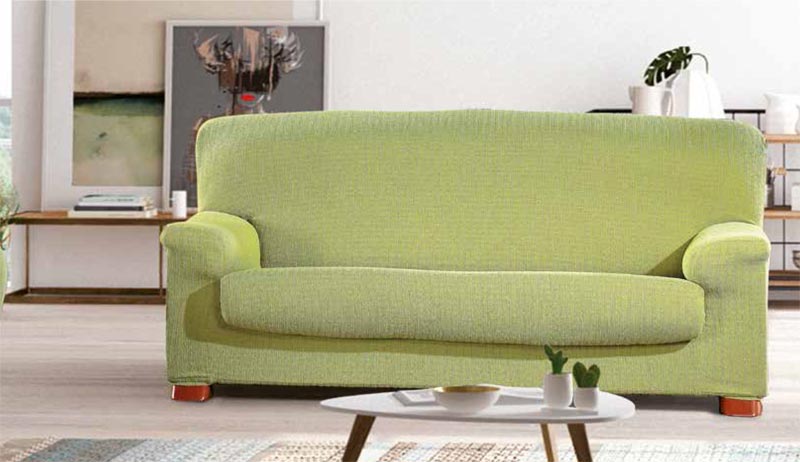
Αγοράστε Στερεό χρώμα αντιολισθητικό κάλυμμα καναπέ παχιά μαλακή βελούδινη πετσέτα μαξιλάρι καναπέ για τα έπιπλα διακόσμησης καθιστικού Slipcovers καλύμματα καναπέ καλύμματα καναπέ | Joom
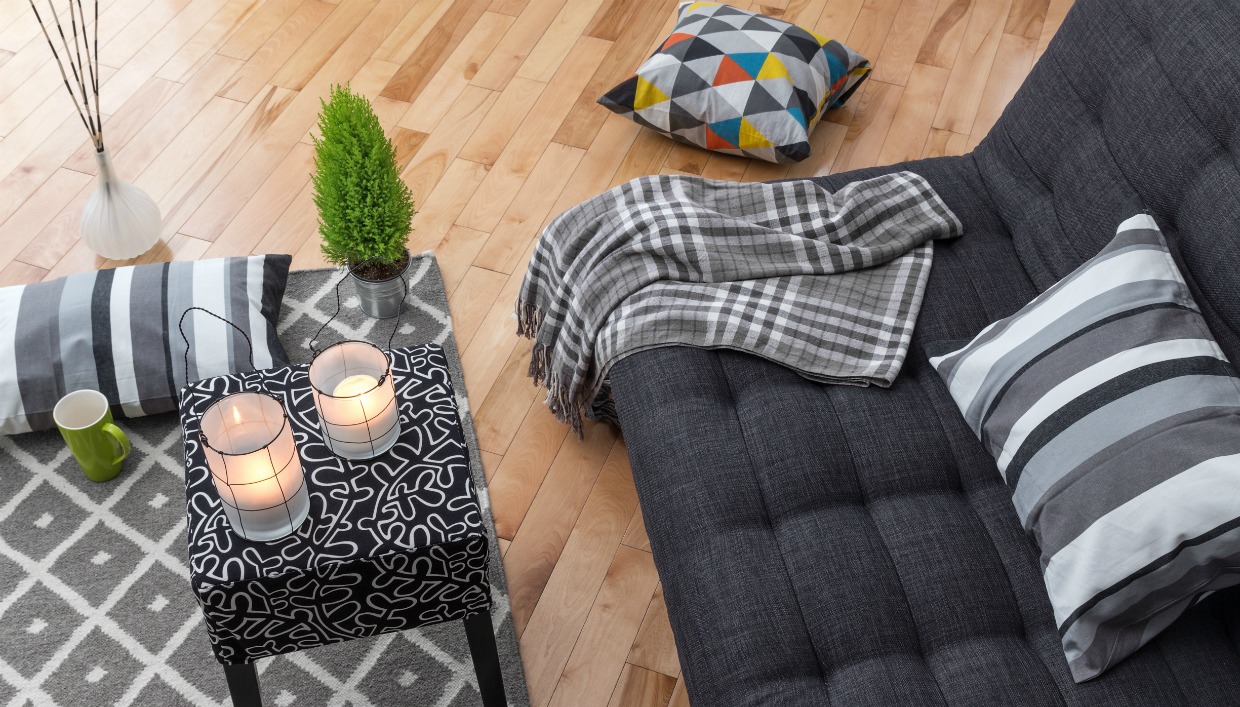

Φτιάξτε Υπέροχα Καλύμματα για τα Μαξιλάρια σας Χωρίς Ράψιμο! - spirossoulis.com - the home issuespirossoulis.com – the home issue

Αγοράστε Στερεό χρώμα αντιολισθητικό κάλυμμα καναπέ παχιά μαλακή βελούδινη πετσέτα μαξιλάρι καναπέ για τα έπιπλα διακόσμησης καθιστικού Slipcovers καλύμματα καναπέ καλύμματα καναπέ | Joom

Αγοράστε Στερεό χρώμα αντιολισθητικό κάλυμμα καναπέ παχιά μαλακή βελούδινη πετσέτα μαξιλάρι καναπέ για τα έπιπλα διακόσμησης καθιστικού Slipcovers καλύμματα καναπέ καλύμματα καναπέ | Joom

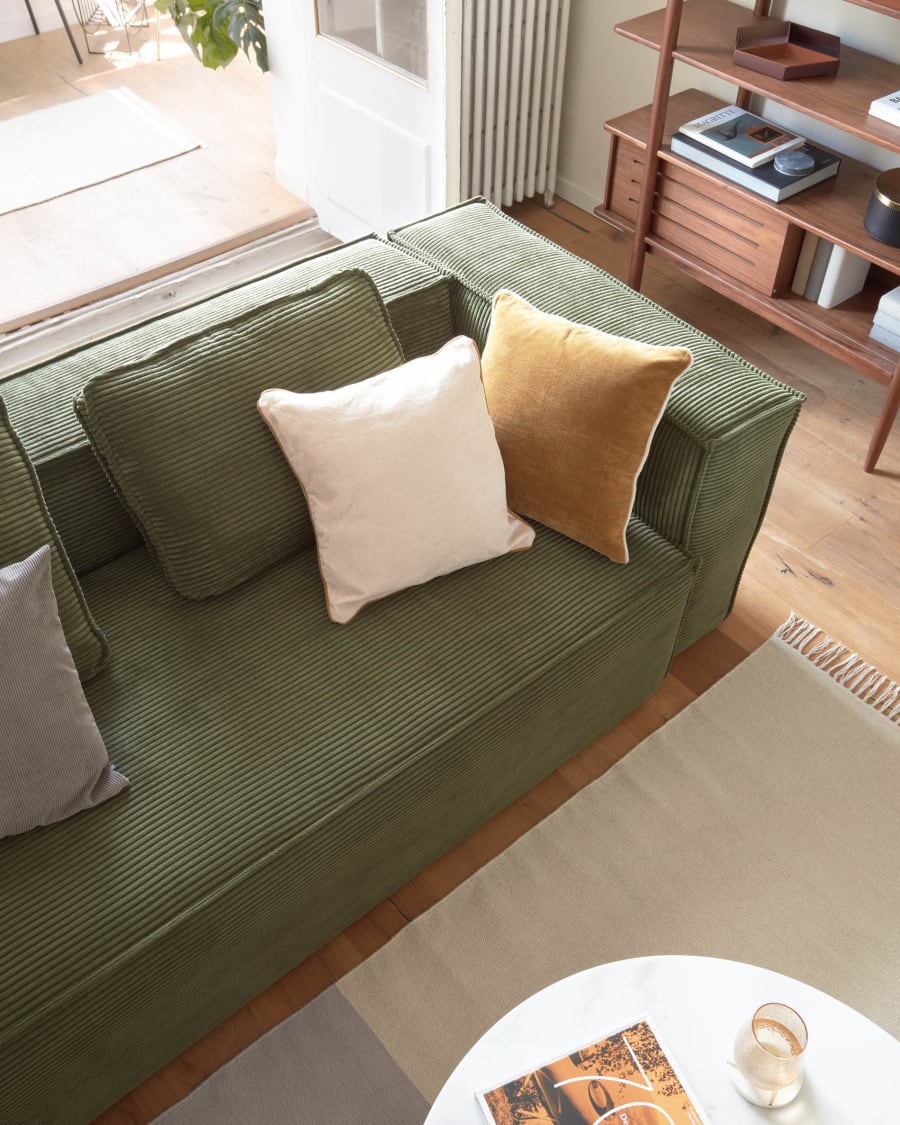
Ελαστικά Καλύμματα Καναπέ Chesterfield Ξεχωριστό Μαξιλάρι Alaska in 2023 | Furniture, Home decor, Love seat